One of the most promising application for deep learning image classification is within the medical field. While imaging technology has allowed doctors to “see” more than ever with a plethora of tests, the diagnosis is still the responsibility of the provider. For example, something simple like an x-ray can reveal many illness about a patient, but it is up to the doctor to interpret the image. This reliance creates an opportunity for human error. To minimize this error, doctors must train extensively to gain both knowledge and experience, but due to fatigue and stress, human error will always exist.
Within the last decade, huge advancements in deep learning and AI have culminated in big projects, such as IBM’s Watson, that have the potential to revolutionize industries. Natural Language Processing (NLP’s) has made it possible to extract data from everyday text data, and image classification is being used by both social media companies and state-of-the-art surveillance systems. Regardless of industry, data science has and will continue to make a big impact.
The goal of this particular project is to employ some common data science techniques in efficiently building a proper convoluted neural network to “read” X-ray scans of pneumonia. The dataset is from kaggle and contains over 6000 images of normal x-rays along with bacterial and viral pneumonia infections. The data is divided into simply normal and pneumonia group, and many kernels on kaggle have attempted binary classification with much success, but few attempted classification between 3 classes. Therefore, the first step is to create new directories and transfer the correct files into them. Luckily, each image is already labeled, so this was not a hard task:

This also meant that the train, test, and validation folder needed to be checked to make sure there is enough data to feed into the model. It turns out that the validation folder had fewer than 10 images, and none of them were viral, so a bit more restructuring was necessary:

With the setup done, the train, validation, and test generators could be set up by the flow_from_directory function. For this project, since there are over 1000 images to train the model on, data augmentation was not used, since that would make things more computationally expensive and extend the run time greatly.

This project includes 6 models, many which used VGG19 as a base model. Each model used slight variations, so to avoid redundancies, only 4 models will be reported. For more information, please see the kaggle link provided. In addition, for the sake of consistency, adam was consistently used as the optimizer and softmax was always the activation function of the final layer of each model.
Model 1: CNN with Alternating Pooling and Convolutional Layers – 75.7% Accuracy

The model had 9 layers and 9 million trainable parameters, which is substantially less than pretrained networks. However, accuracy obtained reached a surprising 75.7%! Using the following function, the training and validation loss and accuracy could also be tracked to check for overfitting via graphing:

The graphs gave no indication of overfitting as both validation and training data followed similar trends:

Model 2: VGG19 without Freezing – 23.8% Accuracy
To import the model, call the function after importing VGG19:
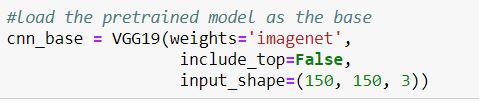
Afterwards, a few dense layers were added to the base for a total of 20 million trainable parameters:

Unfortunately, the model did not achieve a high accuracy at all and scored even below chance level. This suggests that the base layer should be frozen to prevent the weights from being changed by the model.
Model 3: VGG19 with Freezing – 65% Accuracy / 71.5% with best weights
The model is compiled very similarly to before. The only difference is that the trainable attribute of the base model is set to false to prevent the weights from changing:

The model achieved an accuracy of 65%, which is somewhat below the first model. However, this could possibly be improved by loading the best weights saved during the run time:

The accuracy went up a little bit, but not past the first CNN model. While this can suggest that the first model is the best, the relative closeness in accuracy could mean that whatever disparity could be due to the random processes of keras.
Model 4: VGG19 Frozen with Fine-Tuning – 25.6% Accuracy
It is clear that freezing the VGG19 base leads to much better accuracy than letting the model change its weights. On the other hand, fine-tuning can be used to limit how much the model can change the base layer. Ideally, there may be an optimum range for which layers the model should train. The model is set up almost exactly the same as before, but this time, a function is called to unfreeze certain layers within the base CNN. The last 5 layers will be unfrozen for this step:
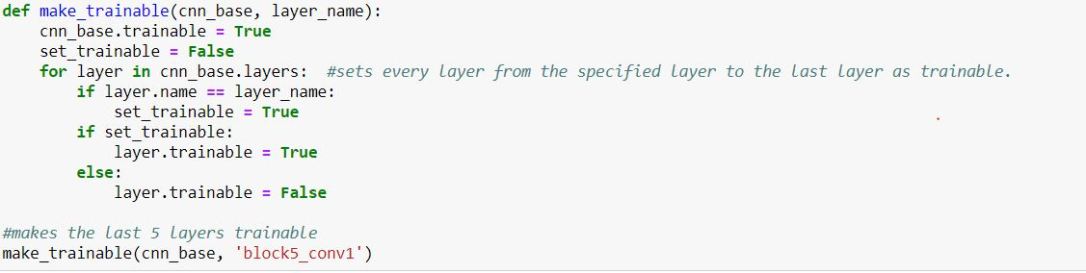
The model achieved only a 25.6% accuracy against the test data. This is negligibly better than allowing the pretained model’s weights to be modified. As a result, this highly suggests that the weights of the pretrained model should not be changed at all, which makes sense due to the fact that VGG19 and other pretrained models were created with much more complex machines and much more data.
Conclusion
It seems like the first model would be the most accurate at 75.7%. However, this is not significantly better than all other models. Because models weights are randomly initialized according to initialization type, it cannot be said with certainty that the first model will always do better. In fact, simply evaluating the model again will lead to a different accuracy score. This is why an ensemble of models are often used instead of just one.What is true is that allowing the pretrained base layer to be changed did lead to a significant decrease in accuracy, as both models’ accuracy dropped below 30%.
While definitely not passable by medical standards, an accuracy rate at about double chance (about 67%) at detecting two different types of pneumonia based solely off of x-ray imaging hints at the power of deep learning. If supplemented by other lab data, it would not be ridiculous to think that machines can be trained to perform better than medical professionals.
